Microsoft Foundry Labs: https://labs.ai.azure.com/projects/gigatime/
GitHub: https://aka.ms/gigatime-code
Sample Data: https://www.dropbox.com/scl/fi/8ampg43fs2yowt9y6vvr1/sample_test_data.zip?rlkey=bkg4w183qnvkh2dudqy3d8lsg&st=j2l463ug&dl=0
HF Model: https://huggingface.co/prov-gigatime/GigaTIME
Listen in multiple languages, when available:
Image Credit: Microsoft Source above
How can AI agents use translators?
Youtube video explains that agents use translators as tools within agentic workflows (1:33).
Specifically:
An assistant (agent) can call the translator, run stratification, draft a report, and then ask for human signoff (1:35-1:43).
Tool-using agents can chain together actions: they can retrieve slides, translate them to virtual channels, compute spatial features, match patient cohorts, and then generate clinician-friendly narratives
Executive Summary
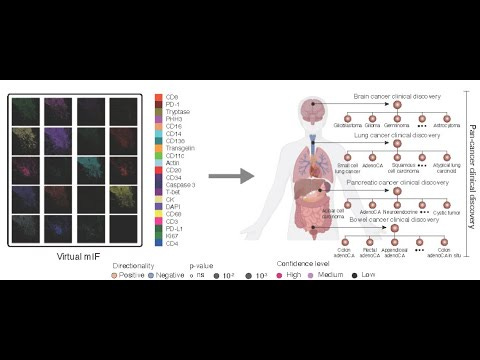
- The paper introduces GigaTIME, a multimodal AI model that translates routinely available H&E pathology slides into high-resolution virtual mIF images to enable population-scale tumor microenvironment (TIME) modeling.
- Trained on a Providence dataset of 40 million cells with paired H&E and mIF across 21 protein channels, GigaTIME bridges cell morphology and cell states, moving beyond prior approaches limited to “average biomarker status.”
- Applied to 14,256 cancer patients across the Providence system, GigaTIME generated a virtual population of ~300,000 mIF images spanning 24 cancer types and 306 subtypes, enabling large-scale discovery of TIME protein–biomarker, staging, and survival associations.
- The study reports 1,234 statistically significant associations linking virtual mIF protein activations to key clinical attributes and demonstrates external validation on 10,200 TCGA patients with strong concordance.
- Compared to CycleGAN baselines, GigaTIME’s cross-modal translation showed superior performance (Dice score and Pearson correlation), supported by qualitative and quantitative analyses.
- The virtual population reveals spatial and combinatorial interactions (entropy, SNR, sharpness; logical OR combinations) that strengthen biomarker associations and improve patient stratification using combined signatures.
- GigaTIME is open-sourced via Microsoft Foundry Labs and Hugging Face, positioning it as a scalable framework for precision immuno-oncology research.
- The work is framed as a step toward a “virtual patient” for high-fidelity digital twins, aligning with Microsoft’s broader multimodal AI efforts in precision health.
What Problem This Solves
- Current mIF spatial proteomics is costly and limited in scalability (“thousands of dollars” per sample), preventing population-scale TIME studies.
- H&E slides are cheap ($5–$10) and routinely available but traditionally used for coarse, average-level biomarker prediction.
- Precision immunotherapy needs spatially resolved, single-cell immune state signals to understand “cold” vs “hot” tumors and predict/improve treatment response.
- GigaTIME exploits multimodal AI to generate virtual mIF from H&E, unlocking large-scale, spatial proteomics-like analyses for:
- Engineering: Scalable cross-modal translation from gigapixel H&E to spatial proteomics signals.
- Product/Clinical research: Population-scale discovery, patient stratification, biomarker association mapping.
- Policy/Health systems: Real-world evidence generation across diverse clinical cohorts and institutions.
Core Contributions (What’s New)
- Introduces a multimodal AI model that “inputs a hematoxylin and eosin (H&E) whole-slide image and outputs multiplex immunofluorescence (mIF) across 21 protein channels.”
- Trains on high-quality paired data of 40 million cells, enabling accurate cross-modal translation beyond prior state-of-the-art (CycleGAN).
- Generates a large virtual mIF population (14,256 patients; ~300,000 images; 24 cancer types; 306 subtypes) for population-scale TIME analysis.
- Discovers 1,234 statistically significant associations linking virtual mIF protein activations to clinical biomarkers, staging, and survival.
- Demonstrates patient stratification improvements using combined multi-channel GigaTIME signatures over individual proteins (e.g., CD3, CD8).
- Reveals spatial and combinatorial interaction metrics (entropy, SNR, sharpness; OR combinations) that enhance biomarker associations.
- Provides independent external validation on 10,200 TCGA patients with strong concordance metrics.
Method / System (How It Works)
Inputs and Outputs
- Inputs:
- H&E whole-slide images (digital pathology) that “reveal details of cell morphology such as nucleus and cytoplasm.”
- Outputs:
- Virtual mIF images across 21 protein channels with spatially resolved single-cell activations.
- Derived “GigaTIME signature” combining all 21 channels for stratification.
Architecture / Components
- Cross-modal translator from digital pathology (H&E) to spatial multiplex proteomics (mIF).
- Compared against CycleGAN for translation performance.
- Not specified in the provided context: precise architectural components (e.g., encoder-decoder details, attention mechanisms specific to GigaTIME), model size, or layers.
Training / Optimization (if described)
- Trained on Providence paired H&E–mIF data covering 40 million cells across 21 protein channels.
- High-quality paired data “enabled much more accurate cross-modal translation compared to prior state-of-the-art methods.”
- Not specified in the provided context: loss functions, optimization algorithms, training schedules, augmentation strategies, or compute resources.
Inference / Runtime (if described)
- Applied inference to generate virtual mIF for 14,256 Providence patients and 10,200 TCGA patients.
- Not specified in the provided context: runtime performance, throughput, hardware requirements, or latency.
Assumptions and Preconditions
- Assumes that H&E-based morphology encodes information about cellular states (“It is well known that H&E-based cell morphology contains information about the cellular states.”).
- Requires availability of high-quality paired H&E–mIF training data for supervised cross-modal learning.
- Assumes generalizability from Providence to external datasets (validated on TCGA).
- Assumes digital pathology slides are routinely available and lower cost vs. mIF, enabling scale.
Evidence and Results (As Reported)
Experimental Setup
- Providence cohort:
- 14,256 patients from 51 hospitals and over a thousand clinics.
- Generated ~300,000 virtual mIF images spanning 24 cancer types and 306 subtypes.
- Training data:
- Paired H&E–mIF covering 40 million cells across 21 protein channels.
- External validation:
- Applied to 10,200 TCGA patients.
- Analyses:
- Correlation analyses for protein–biomarker associations (pan-cancer, type, subtype).
- Survival analyses and patient stratification (e.g., lung, brain; CD3, CD8 vs combined signature).
- Spatial metrics (entropy, SNR, sharpness) and combinatorial channel analyses (logical OR).
- Multiple hypothesis testing correction applied.
Main Results
- Translation performance:
- “Bar plot comparing GigaTIME and CycleGAN on the translation performance in terms of Dice score and Pearson correlation.”
- Qualitative alignment between translated and ground-truth mIF (e.g., DAPI, PD-L1, CD68).
- Population-scale associations:
- Identified “1,234 statistically significant associations between tumor immune cell states (CD138, CD20, CD4) and clinical biomarkers (tumor mutation burden, KRAS, KMT2D).”
- Findings consistent with literature (e.g., “MSI high and TMB high associated with increased activations of TIME-related channels such as CD138.”) and include novel associations (e.g., KMT2D, KRAS).
- Patient stratification:
- “GigaTIME-simulated CD3 and CD8 are similarly effective,” and combining all 21 channels “attained even better patient stratification compared to individual channels.”
- Spatial/combinatorial insights:
- “Associations with spatial features such as sharpness and entropy” and improved biomarker associations via channel combinations (e.g., CD138/CD68; PD-L1/Caspase 3) using logical OR.
- External validation:
- Concordance with TCGA: “Spearman correlation of 0.88 for virtual protein activations across cancer subtypes” and “significant overlap” of associations (“Fisher’s exact test p < 2 × 10−9”).
- Providence yielded “33% more significant associations than TCGA,” highlighting real-world data value.
Ablations / Sensitivity (if described)
- Spatial metrics ablation:
- Entropy, SNR, sharpness compared against activation density to identify protein–biomarker correlations.
- Combinatorial channels:
- Logical OR combinations (e.g., CD138/CD68; PD-L1/Caspase 3) “demonstrating substantially improved associations.”
- Not specified in the provided context: channel-wise or model architecture ablations beyond figures; robustness tests; sensitivity to cohort composition.
Failure Cases (if described)
- Not specified in the provided context.
Key Concepts and Definitions
- H&E (Hematoxylin and Eosin): A common, low-cost stain used in pathology that captures cell morphology.
- mIF (Multiplex Immunofluorescence): Spatial proteomics imaging of multiple protein channels at single-cell resolution; costly and scarce at scale.
- TIME (Tumor Immune Microenvironment): The spatial and functional context of immune cells within tumor tissue, critical for immunotherapy response.
- Virtual population: A large cohort of patients for whom virtual mIF has been generated from H&E slides via AI translation.
- Protein channels: Specific immune/tumor markers (e.g., CD3, CD8, CD138, PD-L1, Caspase 3) measured in mIF.
- GigaTIME signature: Combined signal from all 21 virtual protein channels used for patient stratification.
- OncoTree: A classification system encoding cancer types/subtypes used to organize population-level analyses.
- Real-World Evidence (RWE): Clinical insights derived from real-world patient data across hospitals and clinics.
- Spatial metrics: Quantitative measures of image patterns (entropy, SNR, sharpness) used to capture non-linear spatial interactions.
Implementation Notes (Practical)
- Model access:
- “We have made our GigaTIME model publicly available at Microsoft Foundry Labs and on Hugging Face.”
- Data needs:
- H&E whole-slide images (routine, low-cost, $5–$10 per image).
- Paired H&E–mIF training data improves translation quality (as per Providence dataset).
- Tooling:
- Microsoft Foundry Labs for experimental technologies and potential future integration.
- Deployment constraints:
- Not specified in the provided context: infrastructure requirements, regulatory considerations, PHI handling, inference hardware, throughput, model fine-tuning steps, or data preprocessing pipelines.
Applications in Business
- Precision oncology research:
- Population-scale discovery of TIME protein–biomarker associations to inform clinical hypotheses and trial design.
- Patient stratification:
- Use GigaTIME signatures to stratify patients by staging and survival profiles across cancer types/subtypes.
- Biomarker triaging:
- Complement H&E-based triaging by adding spatially resolved immune state proxies (e.g., CD3, CD8; combined signatures).
- Health system analytics:
- Leverage routinely available H&E slides to generate scalable RWE across large networks (e.g., hospitals and clinics).
- Platform integration:
- Explore integration with advanced multimodal frameworks (e.g., LLaVA-Med) for conversational image analysis.
Risks, Limitations, and Open Questions
- Missing technical specifics:
- Not specified in the provided context: model architecture details, training objectives, compute requirements, and preprocessing.
- Generalization scope:
- External validation on TCGA is reported; broader generalization beyond Providence and TCGA is not specified in the provided context.
- Clinical utility and deployment:
- Not specified in the provided context: prospective clinical trials using virtual mIF for decision-making, regulatory approvals, or workflow integration steps.
- Bias and data representativeness:
- Not specified in the provided context: cohort demographics, site variability control, and potential domain shift handling.
- Measurement fidelity:
- Not specified in the provided context: head-to-head performance against real mIF across clinical endpoints; failure analyses.
Comparative Positioning
- Versus prior H&E-based models:
- Prior works “generally limited to average biomarker status across the entire tissue”; GigaTIME advances to “spatially resolved, single-cell states essential for tumor microenvironment modeling.”
- Versus CycleGAN:
- GigaTIME shows improved translation performance on Dice score and Pearson correlation, supported by bar plots and scatter plots.
- Within Microsoft’s ecosystem:
- Builds on GigaPath for scaling transformers to gigapixel H&E; connects to broader multimodal precision health efforts (BiomedCLIP, LLaVA-Rad, BiomedJourney, BiomedParse, TrialScope, Curiosity).
Actionable Takeaways
- Download and evaluate the open-source GigaTIME model from Foundry Labs or Hugging Face on your existing H&E slide collections.
- Prioritize analyses using combined GigaTIME signatures (all 21 channels) for patient stratification over single-channel approaches.
- Perform correlation studies between virtual mIF activations and available clinical biomarkers; apply multiple hypothesis testing corrections.
- Explore spatial metrics (entropy, SNR, sharpness) and logical OR combinations of channels to strengthen biomarker associations.
- Cross-validate findings against an external cohort (e.g., TCGA) where available to assess concordance.
- Use OncoTree or similar cancer subtype encoding to structure population-scale analyses.
- Document model performance against internal baselines (e.g., CycleGAN or existing workflows) focusing on Dice and Pearson correlations where applicable.
- Plan for data governance around H&E slide ingestion and linkage to clinical attributes; ensure privacy compliance.
- Monitor for domain shifts across sites/hospitals; design validation protocols that reflect clinical diversity.
- Engage clinical stakeholders in interpreting TIME associations for hypothesis generation and trial stratification.
Verbatim Quotes (Evidence)
- “GigaTIME inputs a hematoxylin and eosin (H&E) whole-slide image and outputs multiplex immunofluorescence (mIF) across 21 protein channels.”
- “GigaTIME was trained on a Providence dataset of 40 million cells with paired H&E and mIF images across 21 protein channels.”
- “We applied GigaTIME to 14,256 cancer patients from 51 hospitals and over a thousand clinics within the Providence system.”
- “This effort generated a virtual population of around 300,000 mIF images spanning 24 cancer types and 306 cancer subtypes.”
- “This virtual population uncovered 1,234 statistically significant associations linking mIF protein activations with key clinical attributes such as biomarkers, staging, and patient survival.”
- “Independent external validation on 10,200 Cancer Genome Atlas (TCGA) patients further corroborated our findings.”
- “GigaTIME learned a cross-modal AI translator from digital pathology to spatial multiplex proteomics by training on 40 million cells with paired H&E slides and mIF images from Providence.”
- “The high-quality paired data enabled much more accurate cross-modal translation compared to prior state-of-the-art methods.”
- “We observed significant concordance across the virtual populations from Providence and TCGA, with a Spearman correlation of 0.88 for virtual protein activations across cancer subtypes.”
- “The two populations also uncovered a significant overlap of associations between GigaTIME-simulated protein activations and clinical biomarkers (Fisher’s exact test p < 2 × 10−9).”
- “On the other hand, the Providence virtual population yielded 33% more significant associations than TCGA, highlighting the value of large and diverse real-world data for clinical discovery.”
- “Such a slide only costs $5 to $10 per image and has become routinely available in cancer care.”
One-Page Cheat Sheet
- GigaTIME translates cheap, routine H&E slides into virtual mIF across 21 proteins, enabling spatially resolved single-cell TIME modeling at population scale.
- Trained on 40M cells with paired H&E–mIF; outperforms CycleGAN on Dice and Pearson correlation; qualitative alignment demonstrated.
- Applied to 14,256 Providence patients, generating ~300k virtual mIF images across 24 cancer types and 306 subtypes.
- Discovered 1,234 significant associations between virtual mIF activations and biomarkers, staging, and survival; many align with literature and include novel findings.
- Combined multi-channel “GigaTIME signature” improves patient stratification over single markers (CD3, CD8); supports survival analyses.
- Spatial metrics (entropy, SNR, sharpness) and combinatorial channel ORs (e.g., CD138/CD68; PD-L1/Caspase 3) strengthen biomarker associations.
- External validation on 10,200 TCGA patients shows strong concordance (Spearman 0.88) and significant overlap of associations (Fisher’s p < 2 × 10−9).
- Model available via Microsoft Foundry Labs and Hugging Face; positioned to scale real-world evidence generation in precision oncology.
- Missing details: architecture specifics, training losses, compute, deployment requirements, and prospective clinical utility studies.
- Next steps: deploy open-source model on existing H&E, use combined signatures, analyze biomarker correlations with multiple testing correction, validate across cohorts.
Created with AI